Region of interest pooling explained
Region of interest pooling (also known as RoI pooling) is an operation widely used in object detection tasks using convolutional neural networks. For example, to detect multiple cars and pedestrians in a single image. Its purpose is to perform max pooling on inputs of nonuniform sizes to obtain fixed-size feature maps (e.g. 7×7).
We’ve just released an open-source implementation of RoI pooling layer for TensorFlow (you can find it here). In this post, we’re going to say a few words about this interesting neural network layer. But first, let’s start with some background.
Two major tasks in computer vision are object classification and object detection. In the first case the system is supposed to correctly label the dominant object in an image. In the second case it should provide correct labels and locations for all objects in an image. Of course there are other interesting areas of computer vision, such as image segmentation, but today we’re going to focus on detection. In this task we’re usually supposed to draw bounding boxes around any object from a previously specified set of categories and assign a class to each of them. For example, let’s say we’re developing an algorithm for self-driving cars and we’d like to use a camera to detect other cars, pedestrians, cyclists, etc. — our dataset might look like this.
In this case we’d have to draw a box around every significant object and assign a class to it. This task is more challenging than classification tasks such as MNIST or CIFAR. On each frame of the video, there might be multiple objects, some of them overlapping, some poorly visible or occluded. Moreover, for such an algorithm, performance can be a key issue. In particular for autonomous driving we have to process tens of frames per second.
So how do we solve this problem?
Typical architecture
The object detection architecture we’re going to be talking about today is broken down in two stages:
- Region proposal: Given an input image find all possible places where objects can be located. The output of this stage should be a list of bounding boxes of likely positions of objects. These are often called region proposals or regions of interest. There are quite a few methods for this task, but we’re not going to talk about them in this post.
- Final classification: for every region proposal from the previous stage, decide whether it belongs to one of the target classes or to the background. Here we could use a deep convolutional network.
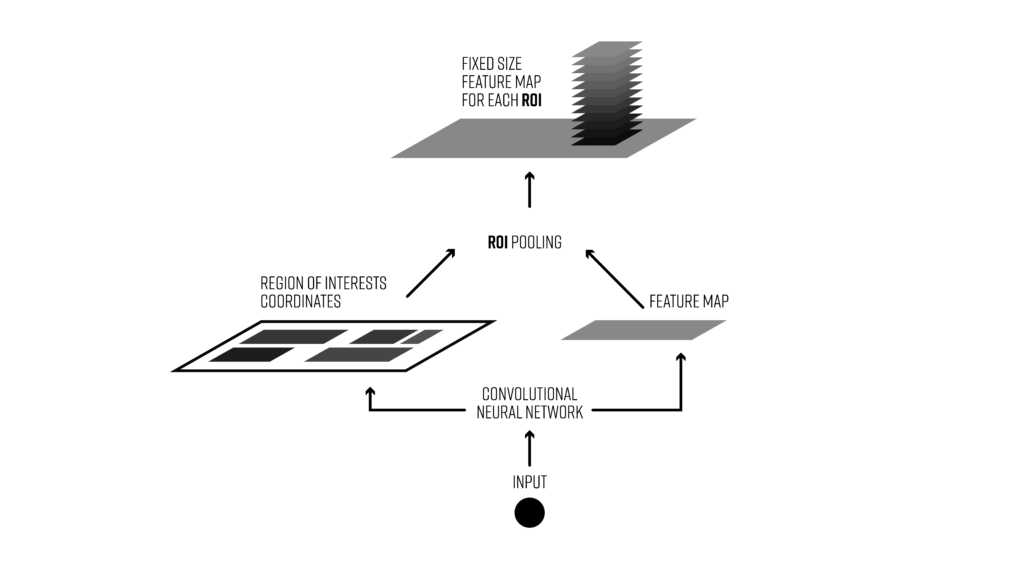
Object detection pipeline with region of interest pooling
Usually in the proposal phase we have to generate a lot of regions of interest. Why? If an object is not detected during the first stage (region proposal), there’s no way to correctly classify it in the second phase. That’s why it’s extremely important for the region proposals to have a high recall. And that’s achieved by generating very large numbers of proposals (e.g., a few thousands per frame). Most of them will be classified as background in the second stage of the detection algorithm.
Some problems with this architecture are:
- Generating a large number of regions of interest can lead to performance problems. This would make real-time object detection difficult to implement.
- It’s suboptimal in terms of processing speed. More on this later.
- You can’t do end-to-end training, i.e., you can’t train all the components of the system in one run (which would yield much better results)
That’s where region of interest pooling comes into play.
What is region of interest pooling?
Region of interest pooling is a neural-net layer used for object detection tasks. It was first proposed by Ross Girshick in April 2015 (the article can be found here) and it achieves a significant speedup of both training and testing. It also maintains a high detection accuracy. The layer takes two inputs:
- A fixed-size feature map obtained from a deep convolutional network with several convolutions and max pooling layers.
- An N x 5 matrix of representing a list of regions of interest, where N is a number of RoIs. The first column represents the image index and the remaining four are the coordinates of the top left and bottom right corners of the region.
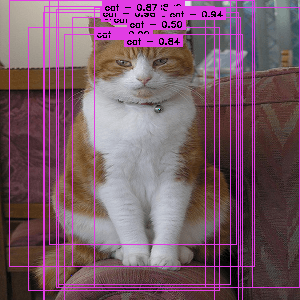 An image from the Pascal VOC dataset annotated with region proposals (the pink rectangles)
An image from the Pascal VOC dataset annotated with region proposals (the pink rectangles)
What does the RoI pooling actually do? For every region of interest from the input list, it takes a section of the input feature map that corresponds to it and scales it to some pre-defined size (e.g., 7×7). The scaling is done by:
- Dividing the region proposal into equal-sized sections (the number of which is the same as the dimension of the output)
- Finding the largest value in each section
- Copying these max values to the output buffer
The result is that from a list of rectangles with different sizes we can quickly get a list of corresponding feature maps with a fixed size. Note that the dimension of the RoI pooling output doesn’t actually depend on the size of the input feature map nor on the size of the region proposals. It’s determined solely by the number of sections we divide the proposal into. What’s the benefit of RoI pooling? One of them is processing speed. If there are multiple object proposals on the frame (and usually there’ll be a lot of them), we can still use the same input feature map for all of them. Since computing the convolutions at early stages of processing is very expensive, this approach can save us a lot of time.
Region of interest pooling — example
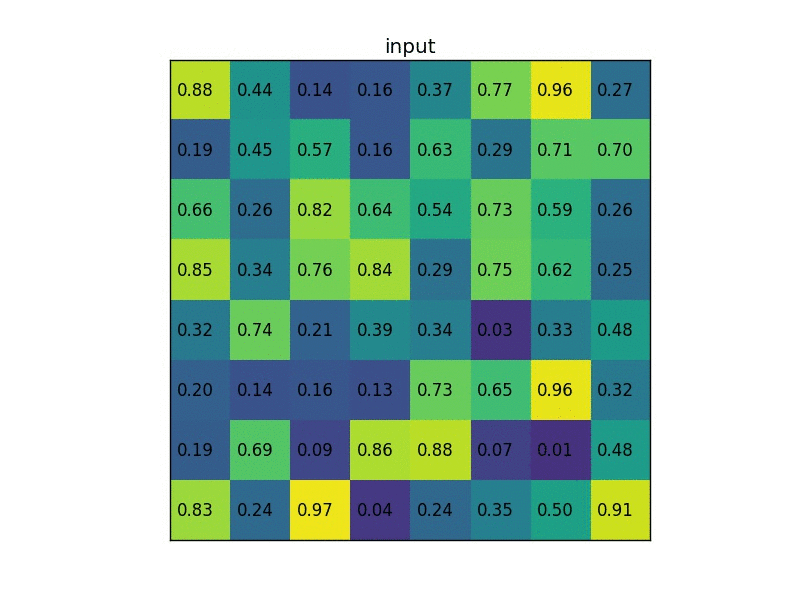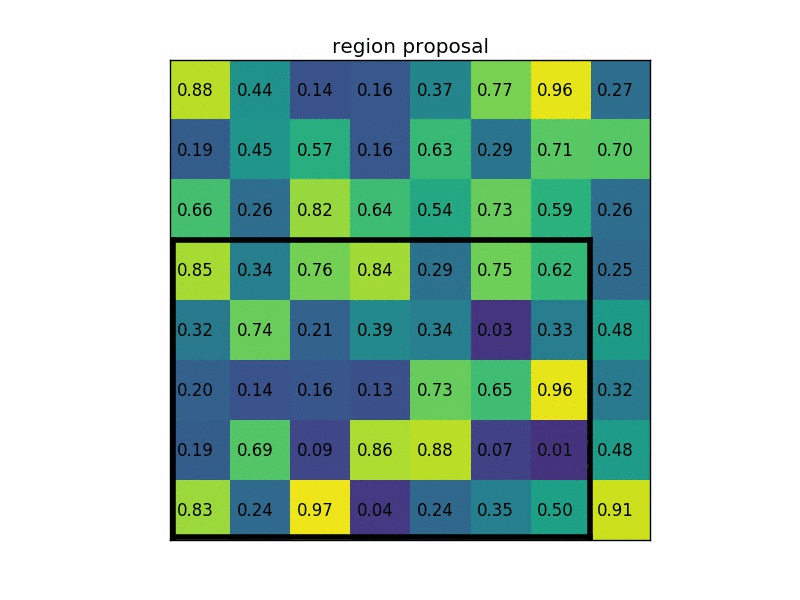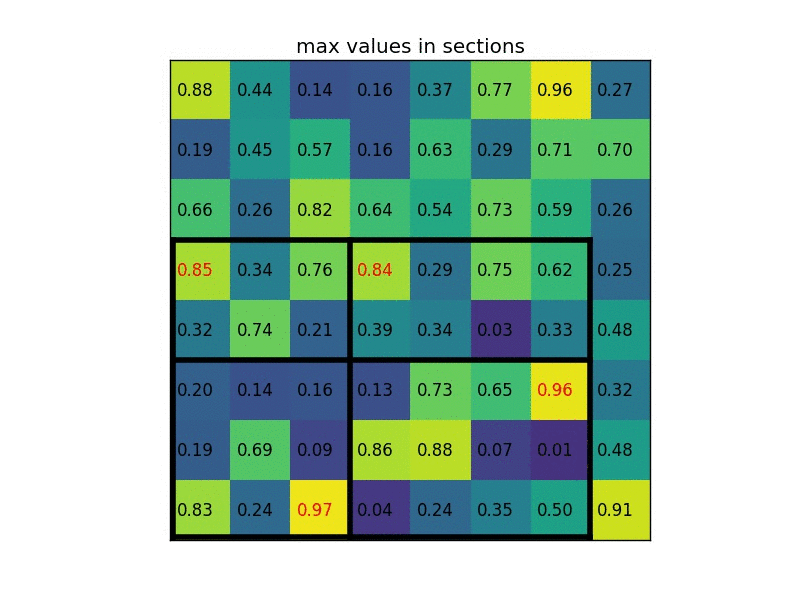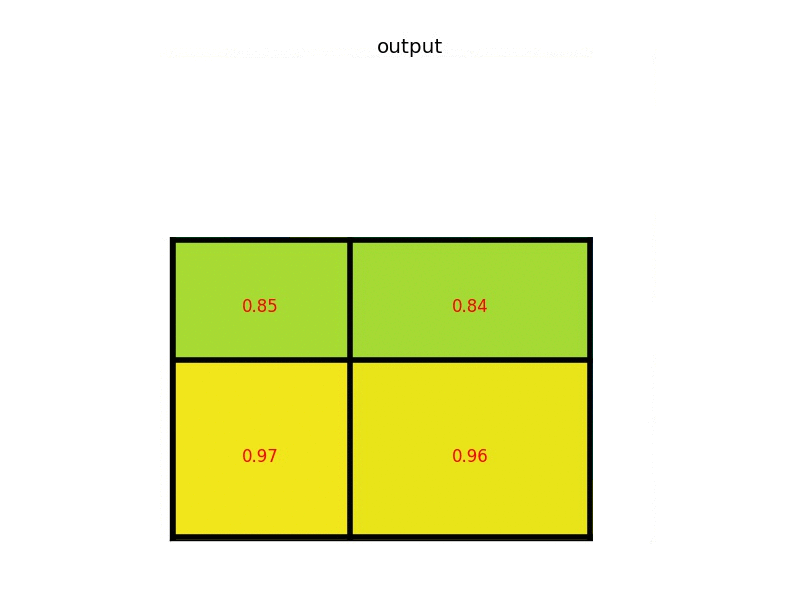
Let’s consider a small example to see how it works. We’re going to perform region of interest pooling on a single 8×8 feature map, one region of interest and an output size of 2×2. Our input feature map looks like this:
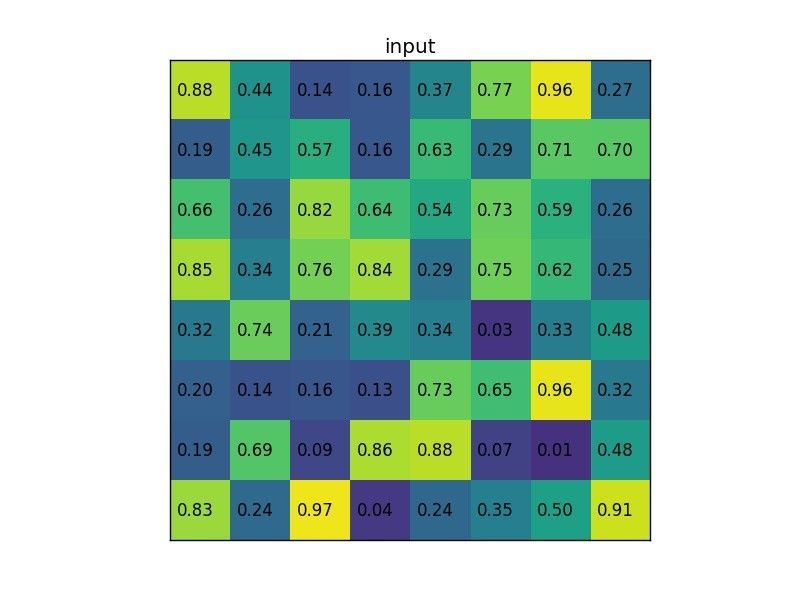
Let’s say we also have a region proposal (top left, bottom right coordinates): (0, 3), (7, 8). In the picture it would look like this:
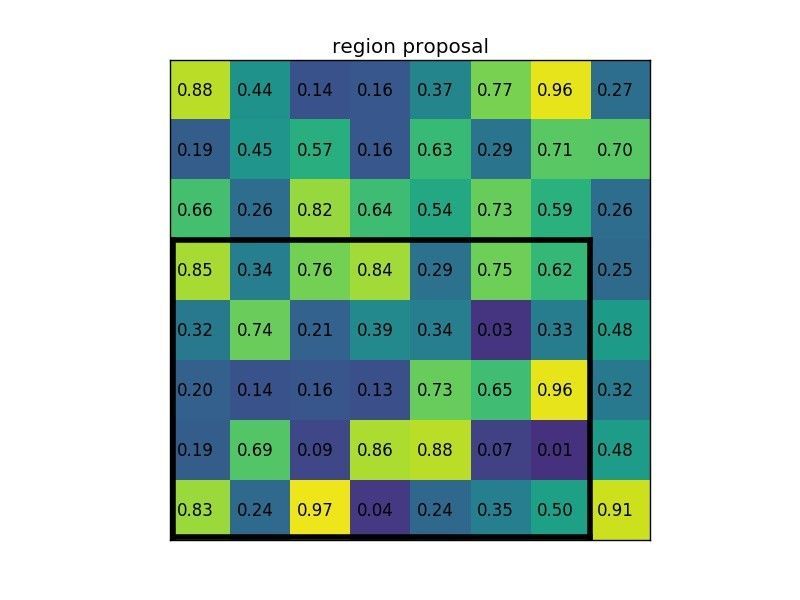 Normally, there’d be multiple feature maps and multiple proposals for each of them, but we’re keeping things simple for the example.
Normally, there’d be multiple feature maps and multiple proposals for each of them, but we’re keeping things simple for the example.
By dividing it into (2×2) sections (because the output size is 2×2) we get:
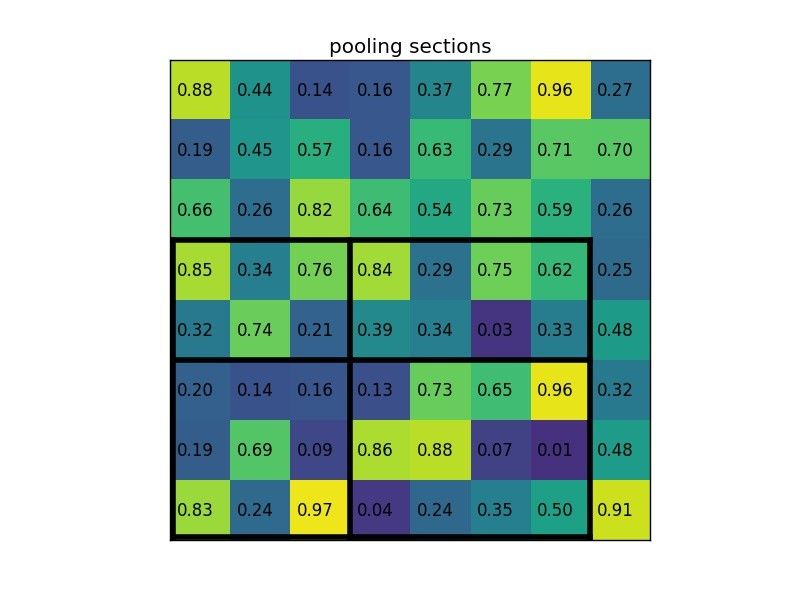
Notice that the size of the region of interest doesn’t have to be perfectly divisible by the number of pooling sections (in this case our RoI is 7×5 and we have 2×2 pooling sections).
The max values in each of the sections are:
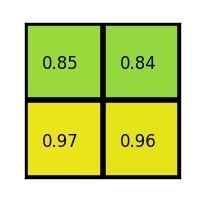
And that’s the output from the Region of Interest pooling layer. Here’s our example presented in form of a nice animation:
What are the most important things to remember about RoI Pooling?
- It’s used for object detection tasks
- It allows us to reuse the feature map from the convolutional network
- It can significantly speed up both train and test time
- It allows to train object detection systems in an end-to-end manner
If you need an opensource implementation of RoI pooling in TensorFlow you can find our version here.
In the next post, we’re going to show you some examples on how to use region of interest pooling with Neptune and TensorFlow.
References
- Girshick, Ross. “Fast r-cnn.” Proceedings of the IEEE International Conference on Computer Vision. 2015.
- Girshick, Ross, et al. “Rich feature hierarchies for accurate object detection and semantic segmentation.” Proceedings of the IEEE conference on computer vision and pattern recognition. 2014.
- Sermanet, Pierre, et al. “Overfeat: Integrated recognition, localization and detection using convolutional networks.” arXiv preprint arXiv:1312.6229. 2013.