Table of contents
Table of contents
In Seahorse we want to provide our users with accurate distributions for their categorical data. Categorical data can be thought of as possible results of an observation that can take one of K possible outcomes. Some examples: Nationality, Marital Status, Gender, Type of Education.
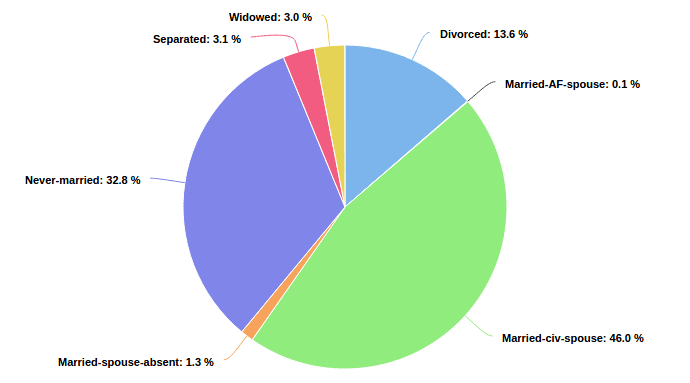 In the described scenario we don’t have prior knowledge whether a given feature is categorical or not. We would like to treat features as categorical as long as their value set is small enough. Moreover, if there are too many distinct values we are no longer interested in its discrete distribution.
Seahorse is build on top of Apache Spark. Naive and easy approach to this problem would be to simply use Spark’s reduceByKey or groupByKey methods. The ReduceByKey method operates on key-value pairs and accepts a function reducing two values into one. If there are many values for one key, those will get reduced to one value with the provided function.
In the described scenario we don’t have prior knowledge whether a given feature is categorical or not. We would like to treat features as categorical as long as their value set is small enough. Moreover, if there are too many distinct values we are no longer interested in its discrete distribution.
Seahorse is build on top of Apache Spark. Naive and easy approach to this problem would be to simply use Spark’s reduceByKey or groupByKey methods. The ReduceByKey method operates on key-value pairs and accepts a function reducing two values into one. If there are many values for one key, those will get reduced to one value with the provided function.
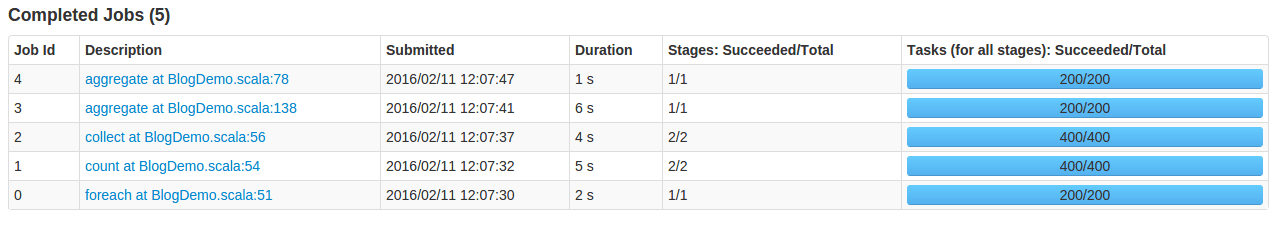 The job 1 (distinct count) and the job 2 (reduceByKey) have 400 tasks and use memory to perform shuffling.
The job 3 and job 4 – have only 200 tasks, run in one stage and don’t use additional memory to perform shuffling. They also check for category counts dynamically.
An interesting observation is that the functional job 3 is slower than the reduceByKey job even though it has only 200 tasks. A memory allocation overhead is that severe!
The job 1 (distinct count) and the job 2 (reduceByKey) have 400 tasks and use memory to perform shuffling.
The job 3 and job 4 – have only 200 tasks, run in one stage and don’t use additional memory to perform shuffling. They also check for category counts dynamically.
An interesting observation is that the functional job 3 is slower than the reduceByKey job even though it has only 200 tasks. A memory allocation overhead is that severe!
 In the described scenario we don’t have prior knowledge whether a given feature is categorical or not. We would like to treat features as categorical as long as their value set is small enough. Moreover, if there are too many distinct values we are no longer interested in its discrete distribution.
Seahorse is build on top of Apache Spark. Naive and easy approach to this problem would be to simply use Spark’s reduceByKey or groupByKey methods. The ReduceByKey method operates on key-value pairs and accepts a function reducing two values into one. If there are many values for one key, those will get reduced to one value with the provided function.
In the described scenario we don’t have prior knowledge whether a given feature is categorical or not. We would like to treat features as categorical as long as their value set is small enough. Moreover, if there are too many distinct values we are no longer interested in its discrete distribution.
Seahorse is build on top of Apache Spark. Naive and easy approach to this problem would be to simply use Spark’s reduceByKey or groupByKey methods. The ReduceByKey method operates on key-value pairs and accepts a function reducing two values into one. If there are many values for one key, those will get reduced to one value with the provided function.
// limit - max. number of distinct categories to treat a feature as categorical
if(rdd.distinct().count() < limit) {
// causes partition reshuffling
val discreteDistribution = rdd.map(v => v -> 1).reduceByKey(_ + _).collect()
Some(discreteDistribution)
} else {
None
}
Unfortunately using these methods means poor performance due to Spark’s partition reshuffling.
Counting occurrences of specific values of a feature can be easily done without reshuffling. We might think of calculating a category distribution as counting occurrences of each category. This can be done using RDD’s function aggregate.
The aggregate function reduces a dataset to one value using a seq and comb functions and an initial aggregator value. The seq function adds one value to an aggregator. It is used to combine all values from a partition into one aggregated value. The comb function adds two aggregators together – it is used to combine results from different partitions.
In our case aggregator value will be of type Option(Map[T, Long]) representing occurrences count for value of type T or None if there are too many distinct categorical values.
The comb function (mergeCombiners in code) adds two aggregator maps together merging their results. If the result map has more distinct categorical values than the specified limit – no distribution is returned (a None).
The seq function (mergeValue in code) increments occurrences count of a value in the aggregator map.
Let the code speak for itself:
import scalaz._
import scalaz.Scalaz._
(...) // other imports
object FunctionalDemo {
// limit - represents the maximum number of categories.
// if there are more distinct values than `limit`, no categorical distribution will be returned
case class CategoryDistributionCalculator[T](limit: Int) {
def calculate(rdd: RDD[T]): Option[Map[T, Long]] =
rdd.aggregate(zero)(mergeValue _, mergeCombiners _)
// initial aggregator value
private val zero = Option(Map.empty[T, Long])
def mergeCombiners[T](
leftOpt: Option[Map[T, Long]],
rightOpt: Option[Map[T, Long]]): Option[Map[T, Long]] = {
for (left <- leftOpt; right <- rightOpt) yield {
// Adds maps in such a way that numbers from same keys are summed.
val sumMap = left |+| right
// yields Some when boolean condition is met. None otherwise
(sumMap.keySet.size <= limit).option(sumMap)
}
}.flatten
def mergeValue[T](acc: Option[Map[T, Long]], next: T): Option[Map[T, Long]] =
mergeCombiners(acc, Some(Map(next -> 1)))
}
def main(args: Array[String]): Unit = {
(...)
val valueSlow = rdd.map(v => v -> 1).reduceByKey(_ + _).collect()
val valueFast = CategoryDistributionCalculator(30).calculate(rdd).get
(...)
}
}
The code above is clean, functional and takes advantage of Scalaz. Unfortunately, it allocates a new object per each record in the dataset. With Apache Spark, sometimes it’s better not to use immutable objects in favor of mutable ones for performance’s sake. According to the documentation, both seq and comb functions are allowed to modify and return their first argument instead of creating a new object to avoid memory allocation. In the next approach we take advantage of that. The code below is not as pretty, but its performance is much better.
(...) // imports
object MutableDemo {
// limit - represents maximum size of categorical type
// if there are more distinct values than `limit`, no categorical distribution will be returned
case class CategoryDistributionCalculator[T](limit: Int) {
// None is returned if there are too many distinct values (>limit) to calculate categorical distribution
def calculate(rdd: RDD[T]): Option[Map[T, Long]] = {
val mutableMapOpt = rdd.aggregate(zero)(mergeValue _, mergeCombiners _)
val immutableMapOpt = mutableMapOpt.map(_.toMap)
immutableMapOpt
}
private val zero = Option(mutable.Map.empty[T, Long])
private def mergeValue(accOpt: Option[mutable.Map[T, Long]],
next: T): Option[mutable.Map[T, Long]] = {
accOpt.foreach { acc =>
addOccurrencesToMap(acc, next, 1)
}
replacedWithNoneIfLimitExceeded(accOpt)
}
private def mergeCombiners(leftOpt: Option[mutable.Map[T, Long]],
rightOpt: Option[mutable.Map[T, Long]]): Option[mutable.Map[T, Long]] = {
for (left <- leftOpt; rightMap <- rightOpt) {
rightMap.foreach { case (element, count) =>
addOccurrencesToMap(left, element, count)
}
}
replacedWithNoneIfLimitExceeded(leftOpt)
}
private def addOccurrencesToMap(
occurrences: mutable.Map[T, Long],
element: T,
count: Long): Unit = {
occurrences(element) = occurrences.getOrElse(element, 0L) + count
}
private def replacedWithNoneIfLimitExceeded(
mapOpt: Option[mutable.Map[T, Long]]): Option[mutable.Map[T, Long]] = {
mapOpt.flatMap { map =>
if (map.size <= limit) mapOpt else None
}
}
}
def main(args: Array[String]): Unit = {
(...)
val valueSlow = rdd.map(v => v -> 1).reduceByKey(_ + _).collect()
val valueFast = CategoryDistributionCalculator(30).calculate(rdd).get
(...)
}
}
Measurements
We put a hypothesis that the last approach should be at least two time faster. The `group by` operation works in two stages with a partition reshuffling between them. Our solution runs in one stage. A simple experiment with Apache Spark seems to confirm that. For the experiment we used:- a local Apache Spark cluster,
- a Data Set containing 10,000,000 rows,
- with 33 labels in the category column (uniformly distributed).
// job 0 - forces Spark to cache data in cluster. It will take away data-loading overhead
// from future jobs and make future measurements reliable.
rdd.foreach((_) => ())
// job 1 - checks if there are too many distinct values to form category
if(rdd.distinct().count() < limit) {
// job 2 - calculates categorical distribution
val discreteDistribution = rdd.map(v => v -> 1).reduceByKey(_ + _).collect()
Some(discreteDistribution)
} else {
None
}
// job 3 - optimised mutable approach
FunctionalDemo.CategoryDistributionCalculator(limit).execute(rdd)
// job 4 - functional approach
MutableDemo.CategoryDistributionCalculator(limit).execute(rdd)
 The job 1 (distinct count) and the job 2 (reduceByKey) have 400 tasks and use memory to perform shuffling.
The job 3 and job 4 – have only 200 tasks, run in one stage and don’t use additional memory to perform shuffling. They also check for category counts dynamically.
An interesting observation is that the functional job 3 is slower than the reduceByKey job even though it has only 200 tasks. A memory allocation overhead is that severe!
The job 1 (distinct count) and the job 2 (reduceByKey) have 400 tasks and use memory to perform shuffling.
The job 3 and job 4 – have only 200 tasks, run in one stage and don’t use additional memory to perform shuffling. They also check for category counts dynamically.
An interesting observation is that the functional job 3 is slower than the reduceByKey job even though it has only 200 tasks. A memory allocation overhead is that severe!